HLA typing by sequence-specific oligonucleotides probes (PCR-SSOP)
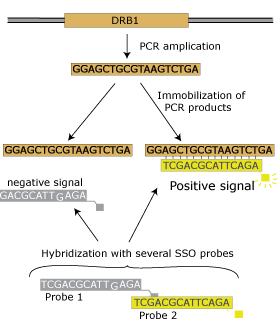
The combination of PCR technology and hybridization with sequence-specific oligonucleotide probes was first applied to HLA class II typing because of the limitations of DR serology and of the better knowledge of allelic polymorphism at DR/DQ loci. The techniqueThe procedure relies on the locus-specific amplification of the genomic DNA segment comprising the polymorphic sites of HLA alleles. Amplified DNA is then immobilized on a solid support, usually a nylon membrane, and then hybridized with a battery of sequence-specific oligonucleotide probes (SSOP) (direct hybridization). Fluorochromes are linked with the probes to allow their detection by chemiluminescence. Alternatively, SSO probes can be immobilized on a solid support, for exemple color-coded microspheres, and hybridized with labeled PCR product (reverse hybridization). The higher the number of probes the better the resolution level. Usually 50-100 probes per locus are used for intermediate/high resolution typing, however, only the probe completely matched with the target sequence amplified will hybridize and give a positive signal. |
|
|
Transplantation Immunology | |||||||||||||||||||
|
||||||||||||||||||||

 Print
Print